-
α-突触核蛋白(α-synuclein, α-syn)在帕金森病的发病机理中起着关键的作用, 因而近年来受到了越来越广泛的关注. α-syn在膜上的动力学过程对理解其功能至关重要. 本文使用基于脂质体的单分子荧光衰减方法—LipoFRET, 首次对较高浓度下α-syn与磷脂膜的相互作用的动态过程进行了单分子层面上的研究. 研究发现, 在溶液中α-syn浓度升高时, 其中央NAC区域可离开磷脂膜表面进入水相中; 而N端部分位于膜表面内的位置变浅, 并有更高的概率脱离膜表面进入溶液中. 利用单分子荧光成像对α-syn解离的观察则发现, 随着溶液中的α-syn浓度升高, 脂质体上的α-syn解离速率加快. 因此高浓度下, α-syn在膜上各区域垂直位置变化促进了蛋白从膜上的解离. 结合LipoFRET的实验结果可以推断, α-syn的解离可能是由于不同的α-syn分子膜作用位点互相竞争而导致的解离. 这样的特征, 可能是体内环境中影响α-syn控制其聚集的重要性质.α-synuclein (α-syn) is a key protein involved in Parkinson’s disease. There have been many researches about α-syn in recent years. It was suggested that the aggregation of α-syn may induce the lipid membranes to disrupted, which is related to the pathology of neurodegenerative diseases. Thus the studying of the dynamics of α-syn on membranes, especially in the presence of high-concentration protein, is important for understanding its function and its role in the pathology. In this study, we use LipoFRET, a single molecule method based on the principle of energy transfer between the donor labeled on the biomolecule and the quenchers encapsulated in the liposome. The quenchers encapsulated in liposomes attenuate the fluorescence attached to membrane proteins near the membrane, or penetrating in the membranes. If interesting site of membrane protein can be labeled, the LipoFRET could probe positional changes of a single membrane protein in the direction normal to the membrane. In the research of α-syn by LipoFRET, some interesting results can be obtained with different concentrations of protein. On the one hand, with the increase of concentration of α-syn in solution, the centre domain of α-syn can leave the surface of the lipid bilayer and enter into the aqueous solution. However, this domain of α-syn is located around the membrane surface at low concentration. On the other hand, the N-terminus of α-syn with three main positions at low concentration of protein, maintains three but different positions in the membrane at high concentration, where each position is closer to or above the outer surface of liposome. The above phenomena suggeste that the interaction between α-syn and membranes might be weakened with the increase of concentration of protein. At the same time, with single molecule fluorescence imaging, we also observe the promoted dissociation rates for individual fluorophore labeled α-syn from liposomes with high concentration of unlabeled proteins in solution. The result is consistent with the result of our single-molecule experiment with LipoFRET. Along with the results from LipoFRET, it could be indicated that there is a competition process where each α-syn could be occupied by the other one at high protein concentration, which leads to the dissociation. The concentration-dependent dissociation may be the property that regulates the aggregation of α-syn in vivo, which is one of the important factors that influence the pathology of the neurodegenerative diseases.
-
Keywords:
- α-synuclein /
- fluorescence resonance energy transfer /
- dissociation /
- membrane-protein interaction
[1] Fusco G, Chen S W, Williamson P T F, Cascella R, Perni M, Jarvis J A, Cecchi C, Vendruscolo M, Chiti F, Cremades N, Ying L M, Dobson C M, de Simone A 2017 Science 358 1440
 Google Scholar
Google Scholar
[2] Sun J C, Wang L N, Bao H, Premi S, Das U, Chapman E R, Roy S 2019 Proc. Natl. Acad. Sci. USA 116 11113
 Google Scholar
Google Scholar
[3] de Weerdt S 2016 Nature 538 S17
 Google Scholar
Google Scholar
[4] 邹昱, 闾坚强, 张庆文, 倪鑫, 陈佩杰, 钱振宇 2019 重庆医科大学学报 44 473
Zou Y, Lü J Q, Zhang Q W, Ni X, Chen P J, Qian Z Y 2019 Jounral of Chongqing Medical University 44 473
[5] Braak H, del Tredici K, Rub U, de Vos R A, Jansen S E N, Braak E 2003 Neurobiol. Aging 24 197
 Google Scholar
Google Scholar
[6] Burre J 2015 J. Park. Dis. 5 699
[7] Vekrellis K, Rideout H J, Stefanis L 2004 Mol. Neurobiol. 30 1
 Google Scholar
Google Scholar
[8] Ulmer T S, Bax A, Cole N B, Nussbaum R L 2005 J. Biol. Chem. 280 9595
 Google Scholar
Google Scholar
[9] Jao C C, Hegde B G, Chen J, Haworth I S, Langen R 2008 Proc. Natl. Acad. Sci. USA 105 19666
 Google Scholar
Google Scholar
[10] Lowe R, Pountney D L, Jensen P H, Gai W P, Voelcker N H 2004 Protein Sci. 13 3245
[11] Diao J, Burre J, Vivona S, Cipriano D J, Sharma M, Kyoung M, Sudhof T C, Brunger A T 2013 eLife 2 e00592
 Google Scholar
Google Scholar
[12] 张振伟, 张建鹏, 周婷婷, 冯伟华, 焦炳华 2010 生命的化学 30 114
 Google Scholar
Google Scholar
Zhang Z W, Zhang J P, Zhou T T, Feng W H, Jiao B H 2010 Chem. Life 30 114
 Google Scholar
Google Scholar
[13] 白佳 2016 博士学位论文 (武汉: 中国科学院武汉物理与数学研究所)
Bai J 2016 Ph. D. Dissertation (Wuhan: Wuhan Institute of Physics and Mathematics, University of Chinese Academy of Sciences) (in Chinese)
[14] Dikiy I, Eliezer D 2012 Biochim. Biophys. Acta 1818 1013
 Google Scholar
Google Scholar
[15] Ferreon A C, Gambin Y, Lemke E A, Deniz A A 2009 Proc. Natl. Acad. Sci. USA 106 5645
 Google Scholar
Google Scholar
[16] van Rooijen B D, Claessens M M, Subramaniam V 2008 Febs Lett. 582 3788
 Google Scholar
Google Scholar
[17] Jo E, McLaurin J, Yip C M, St George-Hyslop P, Fraser P E 2000 J. Biol. Chem. 275 34328
 Google Scholar
Google Scholar
[18] Pfefferkorn C M, Heinrich F, Sodt A J, Maltsev A S, Pastor R W, Lee J C 2012 Biophys. J. 102 613
 Google Scholar
Google Scholar
[19] Jiang Z P, Hess S K, Heinrich F, Lee J C 2015 J. Phys. Chem. B 119 4812
 Google Scholar
Google Scholar
[20] Hellstrand E, Grey M, Ainalem M L, Ankner J, Forsyth V T, Fragneto G, Haertlein M, Dauvergne M T, Nilsson H, Brundin P, Linse S, Nylander T, Sparr E 2013 Acs Chem. Neurosci. 4 1339
 Google Scholar
Google Scholar
[21] Ma D F, Xu C H, Hou W Q, Zhao C Y, Ma J B, Huang X Y, Jia Q, Ma L, Diao J J, Liu C, Li M, Lu Y 2019 Angew. Chem. Int. Ed. 58 5577
 Google Scholar
Google Scholar
[22] Reynolds N P, Soragni A, Rabe M, Verdes D, Liverani E, Handschin S, Riek R, Seeger S 2011 J. Am. Chem. Soc. 133 19366
 Google Scholar
Google Scholar
[23] Burre J, Sharma M, Sudhof T C 2014 Proc. Natl. Acad. Sci. USA 111 E4274
 Google Scholar
Google Scholar
[24] 汪志鹏, 高歌, 段春礼, 杨慧 2018 中国生物化学与分子生物学报 34 1013
Wang Z P, Gao G, Duan C L, Yang H 2018 Chin. J. Biochem. Mol. Biol. 34 1013
[25] Rabe M, Soragni A, Reynolds N P, Verdes D, Liverani E, Riek R, Seeger S 2013 Acs Chem. Neurosci. 4 408
 Google Scholar
Google Scholar
[26] 马丽, 贺小龙, 李明, 胡书新 2018 物理学报 67 148703
 Google Scholar
Google Scholar
Ma L, He X L, Li M, Hu S X 2018 Acta Phys. Sin. 67 148703
 Google Scholar
Google Scholar
[27] 陈泽, 马建兵, 黄星榞, 贾棋, 徐春华, 张慧东, 陆颖 2018 物理学报 67 118201
 Google Scholar
Google Scholar
Chen Z, Ma J B, Huang X Y, Jia Q, Xu C H, Zhang H D, Lu Y 2018 Acta Phys. Sin. 67 118201
 Google Scholar
Google Scholar
[28] Gennis R B, Cantor C R 1972 Biochemistry 11 2509
 Google Scholar
Google Scholar
[29] Lee J, Lee S, Ragunathan K, Joo C, Ha T, Hohng S 2010 Angew. Chem. Int. Ed. 49 9922
 Google Scholar
Google Scholar
[30] Ha T, Enderle T, Ogletree D F, Chemla D S, Selvin P R, Weiss S 1996 Proc. Natl. Acad. Sci. USA 93 6264
 Google Scholar
Google Scholar
[31] Joo C, McKinney S A, Nakamura M, Rasnik I, Myong S, Ha T 2006 Cell 126 515
 Google Scholar
Google Scholar
[32] Diez M, Zimmermann B, Borsch M, Konig M, Schweinberger E, Steigmiller S, Reuter R, Felekyan S, Kudryavtsev V, Seidel C A, Graber P 2004 Nat. Struct. Mol. Biol. 11 135
 Google Scholar
Google Scholar
[33] Kerssemakers J W, Munteanu E L, Laan L, Noetzel T L, Janson M E, Dogterom M 2006 Nature 442 709
 Google Scholar
Google Scholar
[34] Nagle J F, Tristram-Nagle S 2000 Biochim. Biophys. Acta 1469 159
 Google Scholar
Google Scholar
[35] Kamar R I, Banigan E J, Erbas A, Giuntoli R D, de la Cruz M O, Johnson R C, Marko J F 2017 Proc. Natl. Acad. Sci. USA 114 E3251
 Google Scholar
Google Scholar
[36] Fusco G, Pape T, Stephens A D, Mahou P, Costa A R, Kaminski C F, Schierle G S K, Vendruscolo M, Veglia G, Dobson C M, de Simone A 2016 Nat. Commun. 7 12563
 Google Scholar
Google Scholar
-
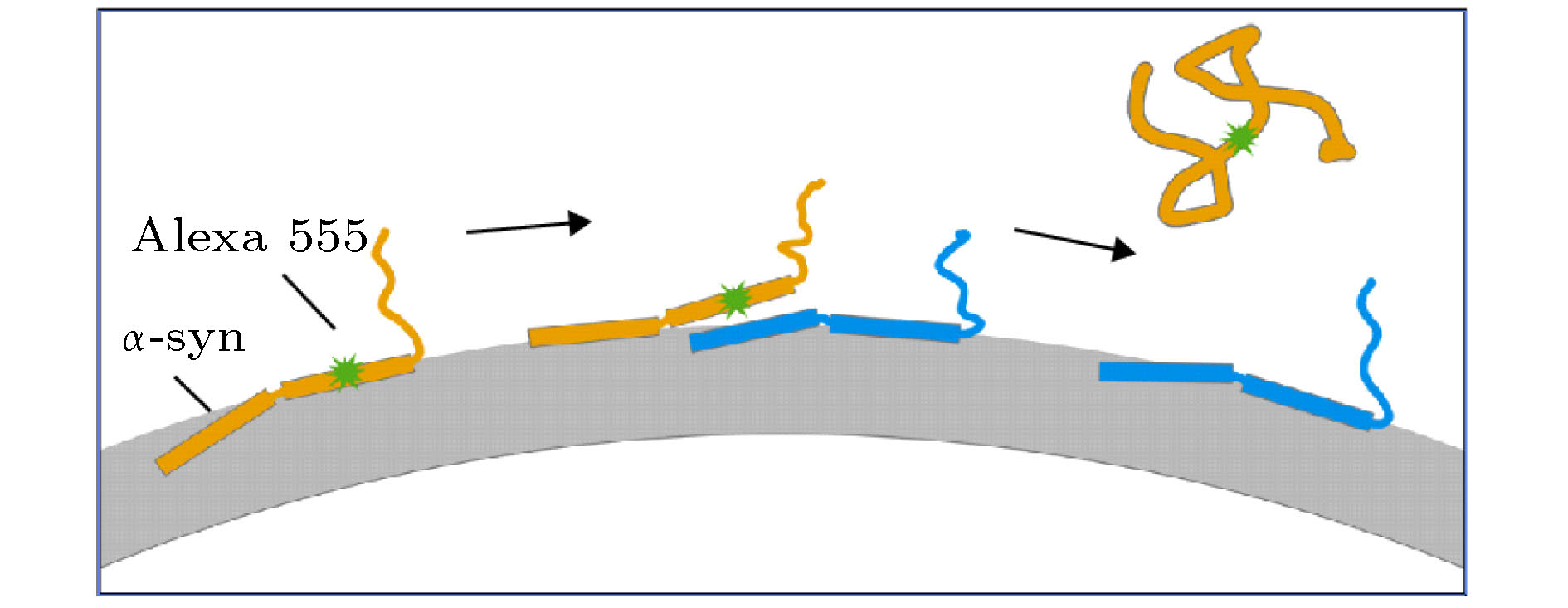
图 1 LipoFRET方法 (a) LipoFRET的实验体系示意图; (b) LipoFRET的距离-相对亮度曲线, 其中淬灭剂为台盼蓝, 所用的荧光分子为Alexa Fluor 555 (Alexa 555), 距离是指荧光基团到脂质体磷脂双层膜的内表面的距离, 曲线分别对应包封2.5 mM (上方曲线)及5 mM (下方曲线)台盼蓝的脂质体.
Fig. 1. The LipoFRET method: (a) The schematic picture of LipoFRET experiment; (b) the relationship between distance from the inner surface of the liposome lipid bilayer and relative intensity F/F0. The quencher used here is trypan blue and the fluorophore is Alexa Fluor 555 (Alexa 555). The curves correspond to liposomes encapsulating 2.5 mM (above) and 5 mM (bottom) trypan blue.
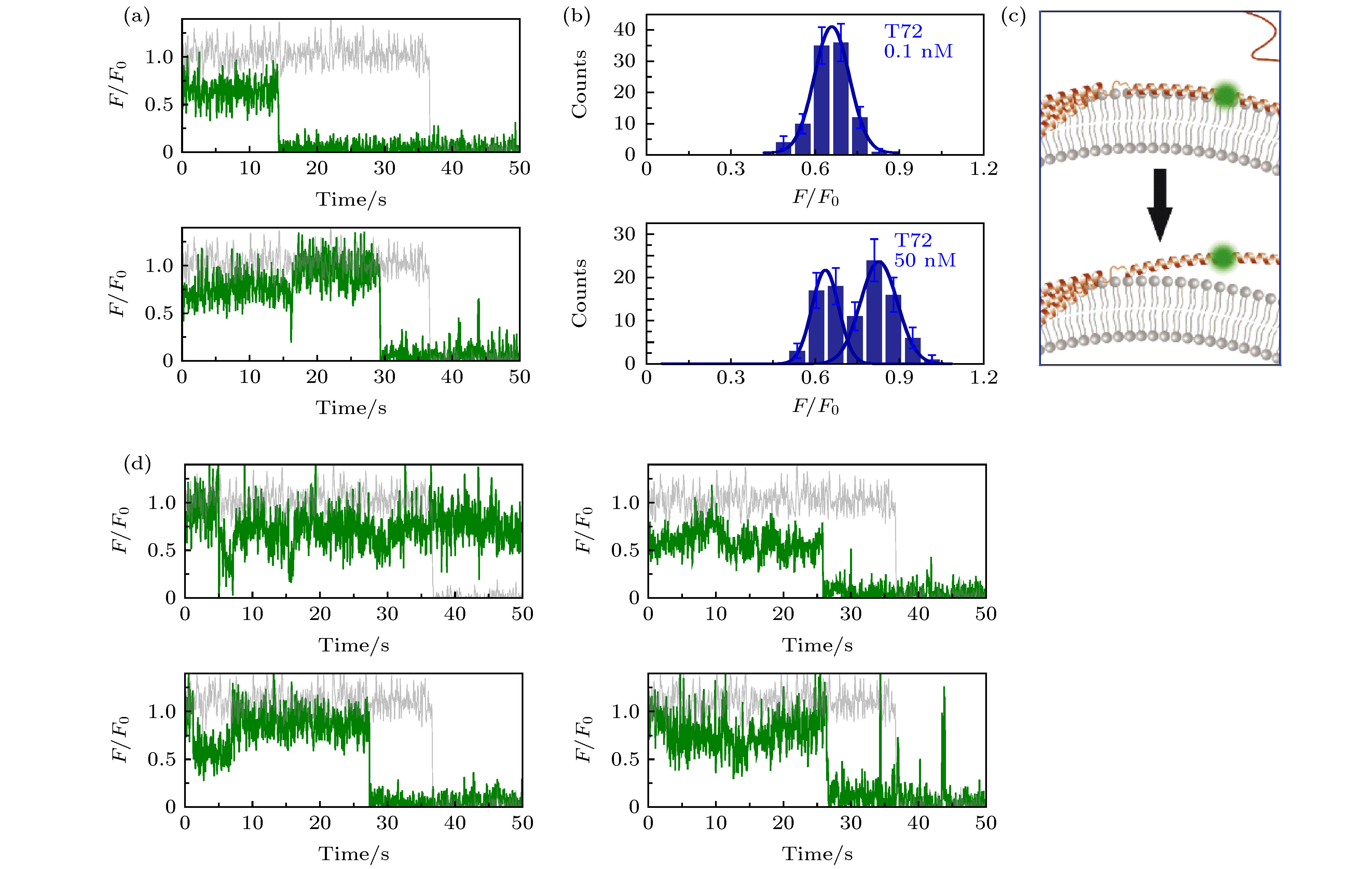
图 2 溶液中存在低蛋白浓度或高蛋白浓度时, α-syn T72C-Alexa 555在包封5 mM台盼蓝的脂质体上的动态特征 (a)蛋白总浓度为0.1 nM (上图)与50 nM (下图)时的典型亮度曲线(其中灰色曲线与绿色曲线分别为包封缓冲液脂质体与包封台盼蓝脂质体上的相对亮度曲线(F/F0)); (b)蛋白总浓度为0.1 nM (上图)与50 nM (下图)时的亮度统计(误差线代表统计偏差); (c)低浓度(上部)与高浓度(下部)时, 在磷脂膜上的位置特征示意图; (d)蛋白总浓度为50 nM时更多的典型亮度曲线
Fig. 2. Dynamic of α-syn T72C-Alexa 555 on liposome encapsulating 5 mM trypan blue in the presence of low and high protein concentration in solution: (a) Typical intensity traces of α-syn T72C-Alexa 555 under 0.1 nM (upper panel) or 50 nM wt-α-syn (lower panel) in solution (the grey and green line correspond to the intensity traces on liposomes entrapping buffer (F0) and on liposomes entrapping trypan blue (F), respectively); (b) the intensity histograms of the states under 0.1 nM (upper panel) or 50 nM wt-α-syn (lower panel) in solution (the error bars on the histograms represent the statistical error in the bins); (c) the scheme of the position change of α-syn T72 on the membrane with increased protein concentration in the solution; (d) more intensity traces of α-syn T72C-Alexa 555 in the presence of 50 nM wt-α-syn in solution.
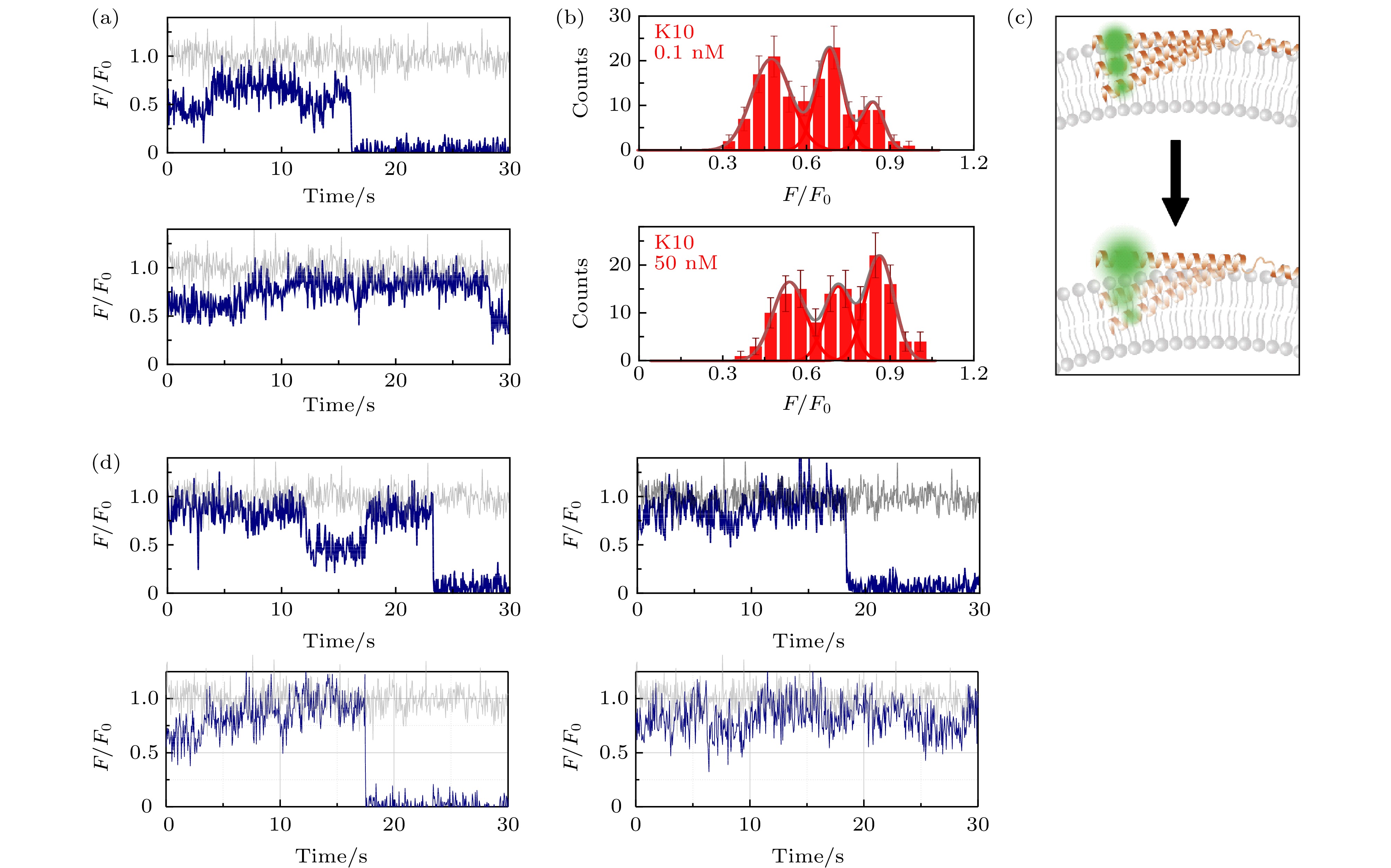
图 3 包封2.5 mM台盼蓝的脂质体上的α-syn K10C-Alexa 555在溶液中存在低浓度与高浓度wt-50 nM wt-α-syn时的动态特征(a)蛋白总浓度为0.1 nM (上图)与50 nM (下图)时的典型亮度曲线(其中灰色曲线与深蓝色曲线分别为包封缓冲液脂质体与包封台盼蓝脂质体上的相对亮度曲线(F/F0)); (b)蛋白总浓度为0.1 nM (上图)与50 nM (下图)时的亮度统计(误差线代表统计偏差); (c)蛋白低浓度(上部)与高浓度(下部)时, 在磷脂膜上的位置特征示意图; (d)蛋白总浓度为50 nM时更多的典型亮度曲线
Fig. 3. Dynamic of α-syn K10C-Alexa 555 on liposome encapsulating 2.5 mM trypan blue in the presence of low and high protein concentration in solution: (a) Typical intensity traces of α-syn K10 C-Alexa 555 under 0.1 nM (upper panel) or 50 nM wt-α-syn (lower panel) in solution (the grey and dark blue line correspond to the intensity traces on liposomes entrapping buffer (F0) and on liposomes entrapping trypan blue (F), respectively); (b) the intensity histograms of the states under 0.1 nM (upper panel) or 50 nM wt-α-syn (lower panel) in solution (the error bars on the histograms represent the statistical error in the bins); (c) the scheme of the position change of α-syn K10 on the membrane with increased protein concentration in the solution; (d) more intensity traces in the presence of 50 nM wt-α-syn in solution.
图 4 溶液中含有1 nM, 10 nM, 100 nM的wt-α-syn时, 视野中脂质体上的α-syn T72C-Alexa 555数量相对于原始数量的比值随时间变化趋势(误差线代表统计偏差)
Fig. 4. Fraction of the remaining fluorescence spots of α-syn T72C-Alexa 555 on the liposomes versus time. The solution contains 1 nM, 10 nM, or 100 nM wt-α-syn. The error bars on the histograms represent the statistical error in the bins.
表 1 α-syn的氨基酸序列
Table 1. Sequences of α-syn.
名称 氨基酸序列(1—140) Human
α-synucleinMDVFMKGLSKAKEGVVAAAEKTKQGVAEAAGKTKEGVL
YVGSKTKEGVVHGVATVAEKTKEQVTNVGGAVVTGVTAV
AQKTVEGAGSIAAATGFVKKDQLGKNEEGAPQEGILEDM
PVDPDNEAYEMPSEEGYQDYEPEA -
[1] Fusco G, Chen S W, Williamson P T F, Cascella R, Perni M, Jarvis J A, Cecchi C, Vendruscolo M, Chiti F, Cremades N, Ying L M, Dobson C M, de Simone A 2017 Science 358 1440
 Google Scholar
Google Scholar
[2] Sun J C, Wang L N, Bao H, Premi S, Das U, Chapman E R, Roy S 2019 Proc. Natl. Acad. Sci. USA 116 11113
 Google Scholar
Google Scholar
[3] de Weerdt S 2016 Nature 538 S17
 Google Scholar
Google Scholar
[4] 邹昱, 闾坚强, 张庆文, 倪鑫, 陈佩杰, 钱振宇 2019 重庆医科大学学报 44 473
Zou Y, Lü J Q, Zhang Q W, Ni X, Chen P J, Qian Z Y 2019 Jounral of Chongqing Medical University 44 473
[5] Braak H, del Tredici K, Rub U, de Vos R A, Jansen S E N, Braak E 2003 Neurobiol. Aging 24 197
 Google Scholar
Google Scholar
[6] Burre J 2015 J. Park. Dis. 5 699
[7] Vekrellis K, Rideout H J, Stefanis L 2004 Mol. Neurobiol. 30 1
 Google Scholar
Google Scholar
[8] Ulmer T S, Bax A, Cole N B, Nussbaum R L 2005 J. Biol. Chem. 280 9595
 Google Scholar
Google Scholar
[9] Jao C C, Hegde B G, Chen J, Haworth I S, Langen R 2008 Proc. Natl. Acad. Sci. USA 105 19666
 Google Scholar
Google Scholar
[10] Lowe R, Pountney D L, Jensen P H, Gai W P, Voelcker N H 2004 Protein Sci. 13 3245
[11] Diao J, Burre J, Vivona S, Cipriano D J, Sharma M, Kyoung M, Sudhof T C, Brunger A T 2013 eLife 2 e00592
 Google Scholar
Google Scholar
[12] 张振伟, 张建鹏, 周婷婷, 冯伟华, 焦炳华 2010 生命的化学 30 114
 Google Scholar
Google Scholar
Zhang Z W, Zhang J P, Zhou T T, Feng W H, Jiao B H 2010 Chem. Life 30 114
 Google Scholar
Google Scholar
[13] 白佳 2016 博士学位论文 (武汉: 中国科学院武汉物理与数学研究所)
Bai J 2016 Ph. D. Dissertation (Wuhan: Wuhan Institute of Physics and Mathematics, University of Chinese Academy of Sciences) (in Chinese)
[14] Dikiy I, Eliezer D 2012 Biochim. Biophys. Acta 1818 1013
 Google Scholar
Google Scholar
[15] Ferreon A C, Gambin Y, Lemke E A, Deniz A A 2009 Proc. Natl. Acad. Sci. USA 106 5645
 Google Scholar
Google Scholar
[16] van Rooijen B D, Claessens M M, Subramaniam V 2008 Febs Lett. 582 3788
 Google Scholar
Google Scholar
[17] Jo E, McLaurin J, Yip C M, St George-Hyslop P, Fraser P E 2000 J. Biol. Chem. 275 34328
 Google Scholar
Google Scholar
[18] Pfefferkorn C M, Heinrich F, Sodt A J, Maltsev A S, Pastor R W, Lee J C 2012 Biophys. J. 102 613
 Google Scholar
Google Scholar
[19] Jiang Z P, Hess S K, Heinrich F, Lee J C 2015 J. Phys. Chem. B 119 4812
 Google Scholar
Google Scholar
[20] Hellstrand E, Grey M, Ainalem M L, Ankner J, Forsyth V T, Fragneto G, Haertlein M, Dauvergne M T, Nilsson H, Brundin P, Linse S, Nylander T, Sparr E 2013 Acs Chem. Neurosci. 4 1339
 Google Scholar
Google Scholar
[21] Ma D F, Xu C H, Hou W Q, Zhao C Y, Ma J B, Huang X Y, Jia Q, Ma L, Diao J J, Liu C, Li M, Lu Y 2019 Angew. Chem. Int. Ed. 58 5577
 Google Scholar
Google Scholar
[22] Reynolds N P, Soragni A, Rabe M, Verdes D, Liverani E, Handschin S, Riek R, Seeger S 2011 J. Am. Chem. Soc. 133 19366
 Google Scholar
Google Scholar
[23] Burre J, Sharma M, Sudhof T C 2014 Proc. Natl. Acad. Sci. USA 111 E4274
 Google Scholar
Google Scholar
[24] 汪志鹏, 高歌, 段春礼, 杨慧 2018 中国生物化学与分子生物学报 34 1013
Wang Z P, Gao G, Duan C L, Yang H 2018 Chin. J. Biochem. Mol. Biol. 34 1013
[25] Rabe M, Soragni A, Reynolds N P, Verdes D, Liverani E, Riek R, Seeger S 2013 Acs Chem. Neurosci. 4 408
 Google Scholar
Google Scholar
[26] 马丽, 贺小龙, 李明, 胡书新 2018 物理学报 67 148703
 Google Scholar
Google Scholar
Ma L, He X L, Li M, Hu S X 2018 Acta Phys. Sin. 67 148703
 Google Scholar
Google Scholar
[27] 陈泽, 马建兵, 黄星榞, 贾棋, 徐春华, 张慧东, 陆颖 2018 物理学报 67 118201
 Google Scholar
Google Scholar
Chen Z, Ma J B, Huang X Y, Jia Q, Xu C H, Zhang H D, Lu Y 2018 Acta Phys. Sin. 67 118201
 Google Scholar
Google Scholar
[28] Gennis R B, Cantor C R 1972 Biochemistry 11 2509
 Google Scholar
Google Scholar
[29] Lee J, Lee S, Ragunathan K, Joo C, Ha T, Hohng S 2010 Angew. Chem. Int. Ed. 49 9922
 Google Scholar
Google Scholar
[30] Ha T, Enderle T, Ogletree D F, Chemla D S, Selvin P R, Weiss S 1996 Proc. Natl. Acad. Sci. USA 93 6264
 Google Scholar
Google Scholar
[31] Joo C, McKinney S A, Nakamura M, Rasnik I, Myong S, Ha T 2006 Cell 126 515
 Google Scholar
Google Scholar
[32] Diez M, Zimmermann B, Borsch M, Konig M, Schweinberger E, Steigmiller S, Reuter R, Felekyan S, Kudryavtsev V, Seidel C A, Graber P 2004 Nat. Struct. Mol. Biol. 11 135
 Google Scholar
Google Scholar
[33] Kerssemakers J W, Munteanu E L, Laan L, Noetzel T L, Janson M E, Dogterom M 2006 Nature 442 709
 Google Scholar
Google Scholar
[34] Nagle J F, Tristram-Nagle S 2000 Biochim. Biophys. Acta 1469 159
 Google Scholar
Google Scholar
[35] Kamar R I, Banigan E J, Erbas A, Giuntoli R D, de la Cruz M O, Johnson R C, Marko J F 2017 Proc. Natl. Acad. Sci. USA 114 E3251
 Google Scholar
Google Scholar
[36] Fusco G, Pape T, Stephens A D, Mahou P, Costa A R, Kaminski C F, Schierle G S K, Vendruscolo M, Veglia G, Dobson C M, de Simone A 2016 Nat. Commun. 7 12563
 Google Scholar
Google Scholar
计量
- 文章访问数: 18889
- PDF下载量: 107
- 被引次数: 0












 下载:
下载: